Most commented posts
- Phonons: shake those atoms — 3 comments
- Start to Fortran — 1 comment
Aug 04 2020
| Authors: | Danny E. P. Vanpoucke, Onno S. J. van Knippenberg, Ko Hermans, Katrien V. Bernaerts, and Siamak Mehrkanoon |
| Journal: | Journal of Applied Physics 128, 054901 (2020) |
| doi: | 10.1063/5.0012285 |
| IF(2019): | 2.286 |
| export: | bibtex |
| pdf: | <JApplPhys> (Open Access) |
| github: | <Amadeus> |
 |
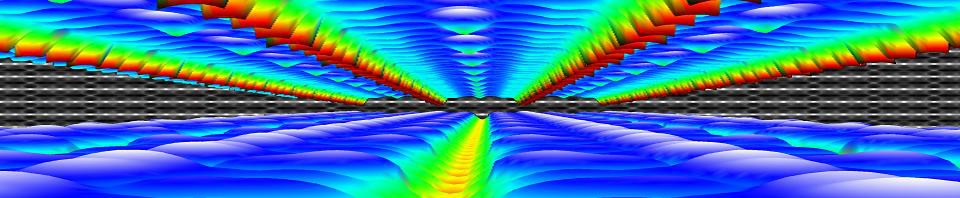
| Graphical Abstract: Correlation plot of the RMSE of the validation set and the intercept value for linear model instances trained on 1000 subsets of a 25 point data set. The distribution of the correlation data is indicated by the black curve. |
Machine Learning is quickly becoming an important tool in modern materials design. Where many of its successes are rooted in huge data sets, the most common applications in academic and industrial materials design deal with data sets of at best a few tens of data points. Harnessing the power of Machine Learning in this context is therefore of considerable importance. In this work, we investigate the intricacies introduced by these small data sets. We show that individual data points introduce a significant chance factor in both model training and quality measurement. This chance factor can be mitigated by the introduction of an ensemble-averaged model. This model presents the highest accuracy while at the same time it is robust with regard to changing data set size. Furthermore, as only a single model instance needs to be stored and evaluated, it provides a highly efficient model for prediction purposes, ideally suited for the practical materials scientist.
Apr 22 2020
| Authors: | Danny E. P. Vanpoucke |
| Journal: | Computational Materials Science 181, 109736 (2020) |
| doi: | 10.1016/j.commatsci.2020.109736 |
| IF(2019): | 2.863 |
| export: | bibtex |
| pdf: | <ComputMaterSci> (Open Access) |
| github: | <Hive-toolbox> |
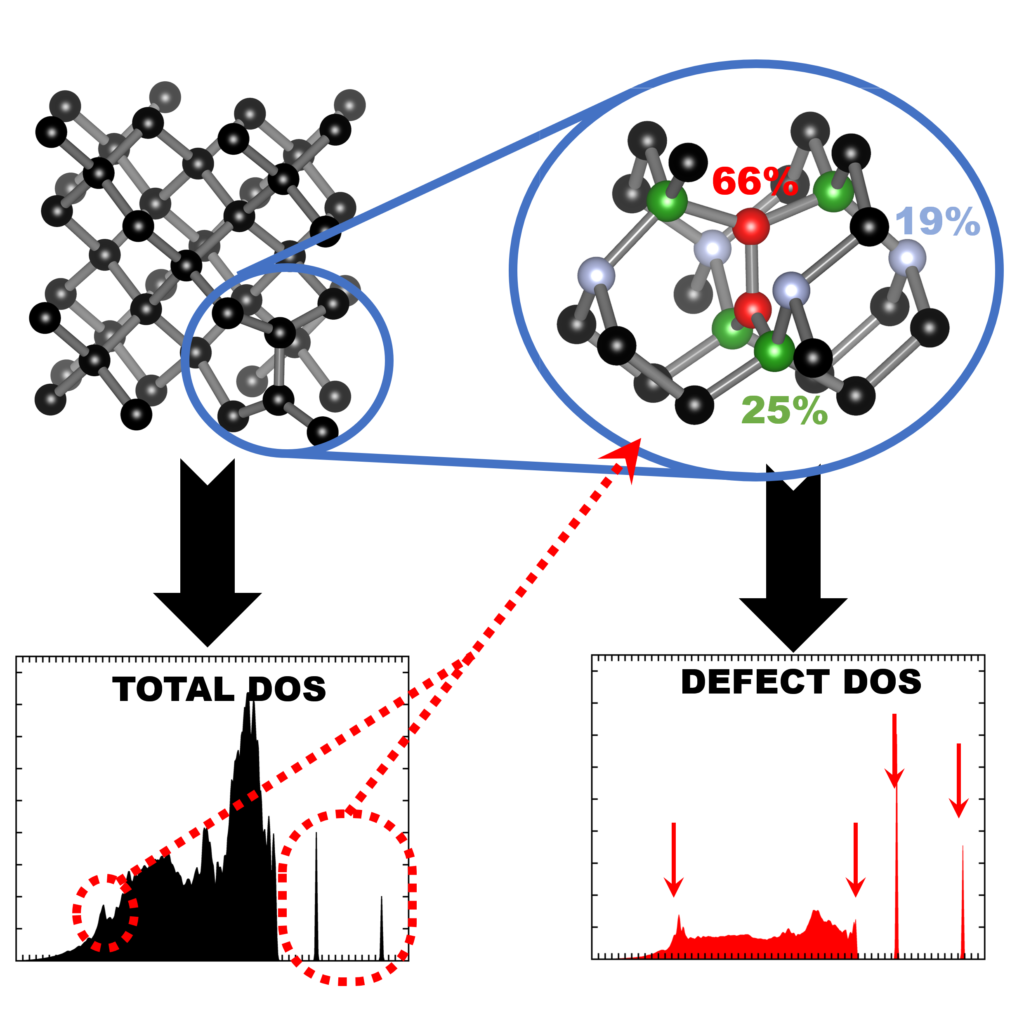 |
| Graphical Abstract: Finger printing defects in diamond through the creation of the vibrational spectrum of a defect. |
Vibrational spectroscopy techniques are some of the most-used tools for materials
characterization. Their simulation is therefore of significant interest, but commonly
performed using low cost approximate computational methods, such as force-fields.
Highly accurate quantum-mechanical methods, on the other hand are generally only used
in the context of molecules or small unit cell solids. For extended solid systems,
such as defects, the computational cost of plane wave based quantum mechanical simulations
remains prohibitive for routine calculations. In this work, we present a computational scheme
for isolating the vibrational spectrum of a defect in a solid. By quantifying the defect character
of the atom-projected vibrational spectra, the contributing atoms are identified and the strength
of their contribution determined. This method could be used to systematically improve phonon
fragment calculations. More interestingly, using the atom-projected vibrational spectra of the
defect atoms directly, it is possible to obtain a well-converged defect spectrum at lower
computational cost, which also incorporates the host-lattice interactions. Using diamond as
the host material, four point-defect test cases, each presenting a distinctly different
vibrational behaviour, are considered: a heavy substitutional dopant (Eu), two intrinsic
point-defects (neutral vacancy and split interstitial), and the negatively charged N-vacancy
center. The heavy dopant and split interstitial present localized modes at low and high
frequencies, respectively, showing little overlap with the host spectrum. In contrast, the
neutral vacancy and the N-vacancy center show a broad contribution to the upper spectral range
of the host spectrum, making them challenging to extract. Independent of the vibrational behaviour,
the main atoms contributing to the defect spectrum can be clearly identified. Recombination of
their atom-projected spectra results in the isolated spectrum of the point-defect.
Mar 24 2020
In the previous sessions of this tutorial on Object Oriented Programming in Fortran 2003, the basics of OO programming, including the implementation of constructors and destructors as well as operator overloading were covered. The resulting classes have already become quite extended (cf. github source). Although at this point it is still very clear what each part does and why certain choices were made, memory fades. One year from now, when you revisit your work, this will no longer be the case. Alternately, when sharing code, you don’t want to have to dig through every line of code to figure out how to use it. These are just some of the reasons why code documentation is important. This is a universal habit of programming which should be adopted irrespective of the programming-language and-paradigm, or size of the code base (yes, even small functions should be documented).
In Fortran, comments can be included in a very simple fashion: everything following the “!” symbol (when not used in a string) is considered a comment, and thus ignored by the compiler. This allows for quick and easy documentation of your code, and can be sufficient for single functions. However, when dealing with larger projects retaining a global overview and keeping track of interdependencies becomes harder. This is where automatic documentation generation software comes into play. These tools parse specifically formatted comments to construct API documentation and user-guides. Over the years, several useful tools have been developed for the Fortran language directly, or as a plugin/extension to a more general tool:
As you can see, there is a lot to choose from, all with their own quirks and features. One unfortunate aspect is the fact that most of these tools use different formatting conventions, so switching from one to the another is not an exercise to perform lightly. In this tutorial, the doxygen tool is used, as it provides a wide range of options, is multi-platform, supports multiple languages and multiple output formats.
As you might already expect, Object Oriented Fortran (f2003) is a bit more complicated to document than procedural Fortran, but with some ingenuity doxygen can be made to provide nice documentation even in this case.
Before you can start you will need to install doxygen:
With a nicely installed doxygen, you can make use of the GUI to setup a configuration suited to your specific needs and generate the documentation for your code automatically. For Object Oriented Fortran there are some specific settings you should consider: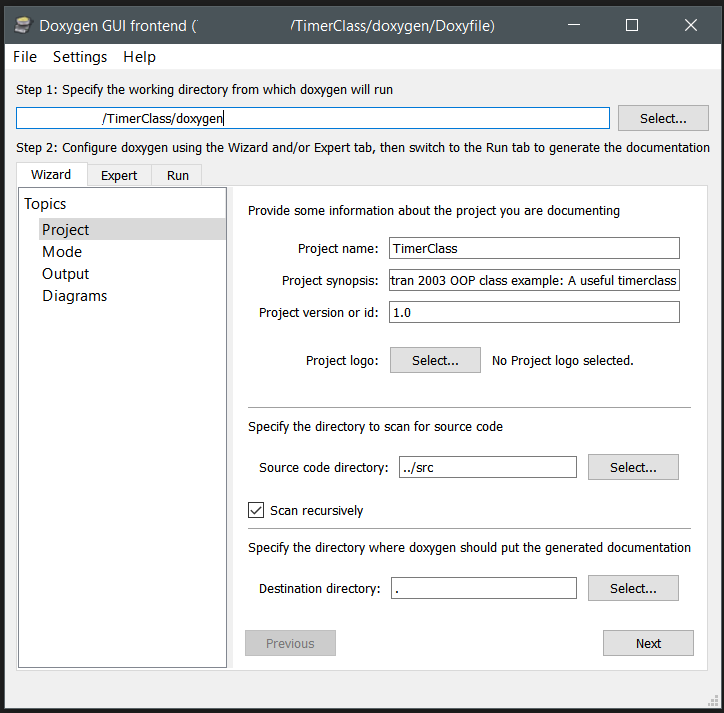
(Provides access each single configuration option to set in doxygen, so I will only highlight a few. Look through them to get a better idea of the capabilities of doxygen.)
Once you have a working configuration for doxygen, you can save this for later use. Doxygen allows you to load an old configuration file and run immediately. The configuration file for the Timer-class project is included in the docs folder, together with the pdf-latex version of the generated documentation. Doxygen generates all latex files required for generating the pdf. To generate the actual pdf, a make.bat file needs to be run (i.e. double-click the file, and watch it run) in a Windows environment.
Let us start with some basics for documenting Fortran code in a way suitable for doxygen. Since doxygen has a very extensive set of options and features, not all of them can be covered. However, the manual of more than 300 pages provides all the information you may need.
With doxygen, you are able to document more or less any part of your code: entire files, modules, functions or variables. In each case, a similar approach can be taken. Let’s consider the documentation of the TimeClass module:
The documentation is placed in a standard single or multi-line fortran comment. In case of multi-line documentation, I have the personal habit turning it into a kind of banner starting with a “!+++++++++” line and closing with a “!<——————-” line. Such choices are your own, and are not necessary for doxygen documentation. For doxygen, a multi-line documentation block starts with “!>” and ends with “!<“ . The documentation lines in between can be indicated with “!!”. This is specifically for fortran documentation in doxygen. C/C++ and other languages will have slightly different conventions, related to their comment section conventions.
In the block above, you immediately see certain words are preceded by an “@”-symbol or a “\”, this indicates these are special keywords. Both the “@” and “\” can be used interchangeably for most keywords, the preference is again personal taste. Furthermore, doxygen supports both html and markdown notation for formatting, providing a lot of flexibility. The multi-line documentation is placed before the object being documented (here an entire module).
Some keywords:
When documenting functions and subroutines there are some addition must-have keywords.
With the knowledge of the previous section, it is relatively easy to document most fortran code. Also the type of object orientation available in fortran 95, in which a fortran module is refurbished as a class. True fortran classes in contrast tend to give a few unexpected issues to deal with. Lets have a look at the documentation of the TTime class of the TimeClass module:
To make sure doxygen generates a class-like documentation for our fortran class, it needs to be told it is a class. This can be done by documenting the class itself and using the keyword @class nameclass, with nameclass the name doxygen will use for this class (so you can choose something different from the actual class name). Unfortunately, doxygen will call this a “module” in the documentation (just poor luck in nomenclature). On the module page for the ttime class a listing is provided of all elements given in the class definition. The documentation added to each member (e.g.,:
is shown as “\brief” documentation. By default all members of our function are considered as public. Adding the @private, @public, or @protected keyword instructs doxygen explicitly to consider these members as private, public or protected. (I used protected in the ttime code not as it should be used in fortran, but as a means of indicating the special status of the final subroutine (i.e. protected in a C++ way).)
However, there seems to be something strange going on. When following the links in the documentation, we do not end up with the documentation provided for the functions/subroutines in the body of our timeclass module. Doxygen seems to consider these two distinct things. The easiest way to link the correct information is by using the keyword @copydoc functionreference . The documentation is (according to doxygen) still for two distinctly different objects, however, this time they have the exact same documentation (unless you add more text on the member documentation line). In this context, it interesting to know there is also @copybrief and @copydetails which can be used to only copy the brief/details section.
In this example, the constructor interface is not documented, as this created confusion in the final documentation since doxygen created a second ttime module/object linked to this interface. However, not documenting this specific instance of the constructor does not create such a large issue, as the module(the fortran module) function itself is documented already.
Documenting fortran classes can be done quite nicely with doxygen. It provides various modes of output: from a fully working website with in-site search engine to a hyperlinked pdf or RTF document. The flexibility and large number of options may be a bit daunting at first, but you can start simple, and work your way up.
As Fortran is supported as an extension, you will need to play around with the various options to find which combination gives the effect you intended. This is an aspect present in all automated code documentation generation tools, since object oriented Fortran is not that widely used. Nonetheless, doxygen provides a very powerful tool worth your time and effort.
Mar 14 2020
Last Wednesday, the 25th edition of the Hasselt Diamond workshop started. The central topic of this celebratory edition was focused on surfaces, perfectly suited to present some of my more recent diamond based work.[1][2] Just as the previous years, the program was packed with interesting talks on anything diamond. Phosphorous doped diamond seemed to be the “new thing” this year, but I could be biased, as I was speaking on phosphorous adsorption myself. Due to a cancellation, I found myself being asked on Monday afternoon to present my work as a talk 😎 , on Wednesday morning 😯 . Because I had been a bit too ambitious in my conference abstract, this talk ended up being nicely complementary to my poster.

Unfortunately, this celebratory edition also fell victim to the COVID-19 crisis. In addition to being the most popular conversation topic—a close second to diamond research—, it also had a very real impact on the conference itself. The COVID-19 crisis resulted in a drop of attendance from 238 people in 2019 to 143 this year. In addition, the quickly changing situation worldwide lead to last minute cancellations due to travel restrictions. On Thursday evening, the conference site went into lock down. Furthermore, that evening, the Belgian federal government also decided that schools and higher education should be closed, as well as pubs and restaurants, until April 3rd. There was also the urgent request for people to work from home as much as possible. (Consider this a good example of acting NOW aimed at saving people.)
Consider this computational scientist in lock down in his home lab until further notice.
Feb 25 2020
| Authors: | Jules Stouten, Danny E. P. Vanpoucke, Guy Van Assche, and Katrien V. Bernaerts |
| Journal: | Macromolecules 53(4), 1388-1404 (2020) |
| doi: | 10.1021/acs.macromol.9b02659 |
| IF(2019): | 5.918 |
| export: | bibtex |
| pdf: | <Macromolecules> (Open Access) |
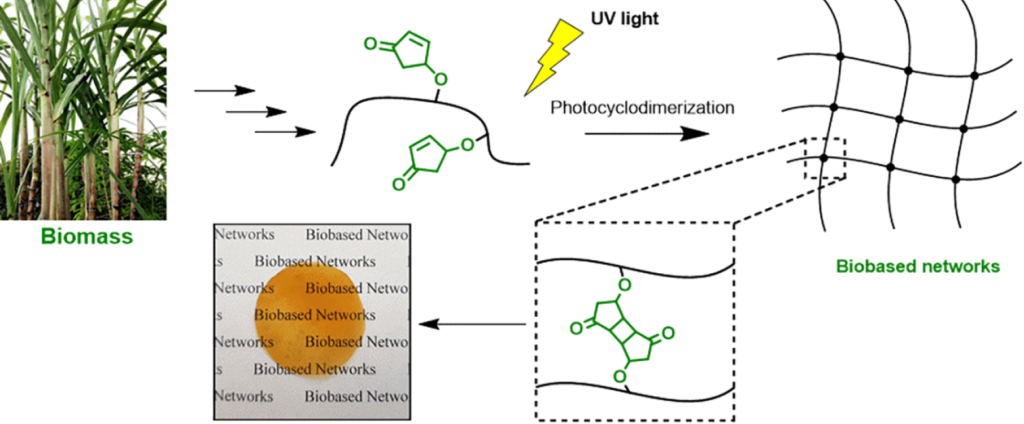 |
| Graphical Abstract: The formation of biobased polyacrylates. |
The controlled polymerization of a new biobased monomer, 4-oxocyclopent-2-en-1-yl acrylate (4CPA), was
established via reversible addition−fragmentation chain transfer (RAFT) (co)polymerization to yield polymers bearing pendent cyclopentenone units. 4CPA contains two reactive functionalities, namely, a vinyl group and an internal double bond, and is an unsymmetrical monomer. Therefore, competition between the internal double bond and the vinyl group eventually leads to gel formation. With RAFT polymerization, when aiming for a degree of polymerization (DP) of 100, maximum 4CPA conversions of the vinyl group between 19.0 and 45.2% were obtained without gel formation or extensive broadening of the dispersity. When the same conditions were applied in the copolymerization of 4CPA with lauryl acrylate (LA), methyl acrylate (MA), and isobornyl acrylate, 4CPA conversions of the vinyl group between 63 and 95% were reached. The additional functionality of 4CPA in copolymers was demonstrated by model studies with 4-oxocyclopent-2-en-1-yl acetate (1), which readily dimerized under UV light via [2 + 2] photocyclodimerization. First-principles quantum mechanical simulations supported the experimental observations made in NMR. Based on the calculated energetics and chemical shifts, a mixture of head-to-head and head-to-tail dimers of (1) were identified. Using the dimerization mechanism, solvent-cast LA and MA copolymers containing 30 mol % 4CPA were cross-linked under UV light to obtain thin films. The cross-linked films were characterized by dynamic scanning calorimetry, dynamic mechanical analysis, IR, and swelling experiments. This is the first case where 4CPA is described as a monomer for functional biobased polymers that can undergo additional UV curing via photodimerization.
Feb 18 2020
| Authors: | Viraj Damle, Kaiqi Wu, Oreste De Luca, Natalia Ortí-Casañ, Neda Norouzi, Aryan Morita, Joop de Vries, Hans Kaper, Inge Zuhorn, Ulrich Eisel, Danny E.P. Vanpoucke, Petra Rudolf, and Romana Schirhagl, |
| Journal: | Carbon 162, 1-12 (2020) |
| doi: | 10.1016/j.carbon.2020.01.115 |
| IF(2019): | 8.821 |
| export: | bibtex |
| pdf: | <Carbon> (Open Access) |
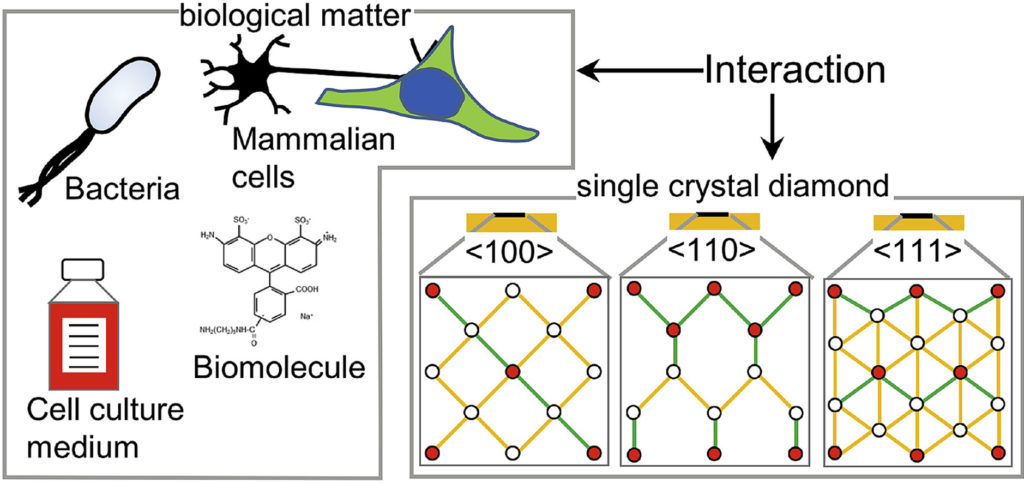 |
| Graphical Abstract: The preferential adsorption of biological matter on oriented diamond surfaces. |
Diamond has been a popular material for a variety of biological applications due to its favorable chemical, optical, mechanical and biocompatible properties. While the lattice orientation of crystalline material is known to alter the interaction between solids and biological materials, the effect of diamond’s crystal orientation on biological applications is completely unknown. Here, we experimentally evaluate the influence of the crystal orientation by investigating the interaction between the <100>, <110> and <111> surfaces of the single crystal diamond with biomolecules, cell culture medium, mammalian cells and bacteria. We show that the crystal orientation significantly alters these biological interactions. Most surprising is the two orders of magnitude difference in the number of bacteria adhering on <111> surface compared to <100> surface when both the surfaces were maintained under the same condition. We also observe differences in how small biomolecules attach to the surfaces. Neurons or HeLa cells on the other hand do not have clear preferences for either of the surfaces. To explain the observed differences, we theoretically estimated the surface charge for these three low index diamond surfaces and followed by the surface composition analysis using x-ray photoelectron spectroscopy (XPS). We conclude that the differences in negative surface charge, atomic composition and functional groups of the different surface orientations lead to significant variations in how the single crystal diamond surface interacts with the studied biological entities.
Jan 01 2020
2019 has come and gone. 2020 eagerly awaits getting acquainted. But first we look back one last time, trying to turn this into a old tradition. What have I done during the last year of some academic merit.
Publications: +3 (and currently +5 submitted)
Completed refereeing tasks: +9
Conferences & workshops: +7 (Attended)
Science Communication Events: +3
Research Stay: +1 With Prof. Klauss-Uwe Koch, Westfälishe Hochschule, Recklinghausen, Germany, July 29th – August 2nd, 2019
PhD-students: +1 Guillaume Emerick (September 2019-August 2023,PhD student UHasselt-UNamur Project, Belgium, Awarded grant for this project)
Bachelor-students: +1 Siebe Frederix (3rd Bach. Phys., Project: Atoms in Molecules based on force partitioning)
Positions: +1 Started working on Machine Learning at AMIBM of Maastricht University
Current size of HIVE:
Hive-STM program:
Dec 27 2019
As part of my machine learning research at AMIBM, I recently ran into the following challenge: “Is it possible to do parallel computation using python.” It sent me on a rather long and arduous journey, with the final answer being something like: “very reluctantly“.
Python was designed with one specific goal in mind; make it easy to implement small test programs to see if an idea is worth pursuing. This gave rise to a scripting language with a lot of flexibility, but also with significant limitations, most of which the “intended” user would never meet. However, as a consequence of its success, many are using it going far beyond this original scope (yours truly as well 🙂 ).
Python offers various libraries to parallelize your scripts…most of them wrappers adding minor additional functionality. However, digging down to the bottom one generally ends up at one of the following two libraries: the threading module and the multiprocessing module.
Of course, as with many things python, there is a huge amount of tutorials available with many of great quality.
Programmers experienced in a programming language such as C/C++, Pascal, or Fortran, may be familiar with the concept of multi-threading. With multi-threading, a CPU allows a program to distribute its work over multiple program-threads which can be performed in parallel by the different cores of the CPU (or while a core is idle, e.g., since a thread is waiting for data to be fetched). One of the most famous API’s for writing multi-threaded applications is OpenMP. In the past I used it to parallelize my Hirshfeld-I implementation and the phonon-module of HIVE.
For Python, there is no implementation of the OpenMP API, instead there is the threading module. This provides access to the creation of multiple threads, each able to perform their own tasks while sharing data-objects. Unfortunately, python has also the Global Interpreter Lock, GIL for short, which allows only a single thread to access the interpreter at a time. This effectively reduces thread-based parallelization to a complex way of running a code in a serial way.
For more information on “multi-threading” in python, you can look into this tutorial.
In addition to the threading module, there is also the multiprocessing module. This module side-steps the GIL by creating multiple processes, each having its own interpreter. This however comes at a cost. Firstly, there is a significant computational cost starting the different processes. Secondly, objects are not shared between processes, so additional work is needed to collect and share data.
Using the “Pool” class, things are somewhat simplified, as can be seen in the code-fragment below. With the pool class one creates a set of threads/processes available for your program. Then through the function apply_async function it is possible to run processes in parallel. (Note that you need to use the “async” version of the function, as otherwise you end up with running things serial …again)
- import multiprocessing as mp
-
- def doOneRun(id:int): #trivial function to run in parallel
- return id**3
-
-
-
- num_workers=10 #number of processes
- NRuns=1000 #number of runs of the function doOneRun
-
- pool=mp.Pool(processes=num_workers) # create a pool of processes
- drones=[pool.apply_async(doOneRun, args=nr) for nr in range(NRuns)] #and run things in parallel
-
- for drone in drones: #and collect the data
- Results.collectData(drone.get()) #Results.collectData is a function you write to recombine the separate results into a single result and is not given here.
-
- pool.close() #close the pool...no new tasks can be run on any of the processes
- pool.join() #collapse all threads back into the main thread
If you are used to HPC applications, you always want to get as much out of your machine as possible. With regard to parallelization this often means making sure no CPU cycle is left unused. In the example above we manually selected the number of processes to spawn. However, would it not be nice if the program itself could just set this value to be equal to the number of physical cores accessible?
Python has a large number of functions claiming to do just that. A few of them are given below.
In conclusion, there seems to be no simple way to obtain the correct number of physical cores using python, and one is forced to provide this number manually. (If you do have knowledge of such a function which works in both windows and unix environments and both desktop and HPC architectures feel free to let me know in the comments.)
All in all, it is technically possible to run code in parallel using python, but you have to deal with a lot of python quirks such as GIL.
Nov 30 2019
Last Tuesday? I had the pleasure of competing in the casting keynotes competition of the TEDx UHasselt chapter. An evening filled with interesting talks on subjects ranging from the FAIR principles of open-data (by Liebet Peeters) to the duty not stay silent in the face of “bad ideas” and leading a life of purpose. An interesting presentation was the one by Ann Bessemans on visual prosody to improve reading skills in young children as well as reading experience, more specifically the transfer of non-literal-content, for non-native speakers. There was also time for some humor, with the dangerous life of Tim Biesmans, who suffers from peanut-allergies. For him, death lurks around every corner, even in a first-date’s kiss. During my talk, I traced the evolution of computational research as the third paradigm of scientific discovery, showing you can find computational research in every field, and why it is evolving at its break-neck speed.
During the event, both the public and a jury voted on the best presentation, which would then have to present at the TEDx UHasselt in 2020.
And the Winner is …drum roll… Danny Vanpoucke!
So this story will continue during the 2020 TEDx event at UHasselt, and I hope to see you there 🙂

top: Full action shots of my presentation. Moore’s Law as driving force behind computational research, and pondering the meaning of Artificial Intelligence. Bottom: Yes, I won 🙂